In previous lectures we described how neural networks can be used to make predictions and classifications. In the classification setting, under the hood it is actually regression being performed and taking a softmax of the outputs in order to return a normalised pseudo-probability of classification for each potential category. That pseudo-probability is very much pseudo-, as that value is relative to other output categories.
Bayesian Neural Networks will change that. And as you will see, Variational Inference (which can be used for Bayesian Neural Networks) can be used to approximate and propagate uncertainty in complex models where it would be too expensive to do so otherwise.
Bayesian Neural Networks
A Bayesian Neural Network can have any architecture that you would otherwise have for a "standard" neural network. The principle difference introduced by the prefix Bayesian is that we will:
- Introduce a prior \(p(w)\) on all the weights \(w\) in the network.
- Propagate our uncertainty in the weights to the output predictions such that we have posteriors of belief for a given class (or posterior predictions).
The claim is that Bayesian Neural Networks are able to capture epistemic uncertainty: describing the uncertainty in having the right model.numpy.Stan or PyMC and just build your neural network with the weights as model parameters and use Hamiltonian Monte Carlo to sample the parameters. In this kind of situation we would expect that the priors on the weights would act to regularise the weights, and (may) avoid us from needing to perform dropout or cross-validation.
Here's what that might look like in Python, where we can feed this model directly to PYMC and use a Hamiltonian Monte Carlo sampler to sample posteriors on the weights in the network:
But if our neural network was very deep, with many tens or hundreds of thousands of weights? Hamiltonian Monte Carlo is efficient, but can we afford to be sampling so many model parameters? And should we explicitly be calculating partial derivatives between all model parameters many parameters are uncorrelated? The answer to all of these questions is no. Thankfully, there are methods of approximating densities — including posterior densities — that we can use. These methods are described within the field of variational inference and are sufficiently powerful such that they can make an intractable problem tractable.stan).
Variational Inference
Imagine you have some complex density that you would like to describe. That density might be a posterior density, and by describing it here I mean we would like to draw samples from that density so you can report quantiles, et cetera. Sampling that density can be extremely expensive, or even intractable.
In variational inference we chose a distribution family that could describe the target density (our posterior) that we do care about, and replace the sampling problem as an optimisation problem. Even though optimisation is Hard™, it is much easier and much more efficient than sampling. So any time you can cast an inference problem as an optimisation problem, you are replacing a very hard problem with a (relatively) easy problem! There are a few steps to understand how variational inference is implemented in practice in order to make it generalisable. I recommend this lecture by David Blei (Columbia) if you are interested.
Suppose our intractable probability distribution is \(p\). Variational inference will try to solve an optimisation problem over a class of tractable distributions \(\mathcal{Q}\) in order to find a \(q \in \mathcal{Q}\) that is most similar to \(p\), where similarity is measured by the divergence
We need some measure for how close \(q\) and \(p\) need to be, and what exactly close means here. In most variational inference problems we use the Kullback-Leibler (K-L) divergence, because it makes the optimisation problem tractable.
Consider then if we have some intractable posterior distribution \(p\) and we are going to approximate it with some distribution \(q\). All we would need to do is minimize the K-L divergence from \(p\) to \(q\). It turns out optimising the K-L divergence \(\mathcal{KL}(q\,||\,p)\) is not possible because of the intractable normalisation constant (the fully marginalised likelihood) in our posterior \(p\). In fact, we can't even properly evaluate \(p\) because we don't know the normalisation constant. Instead, we optimise something that involves the unnormalised proabability, known as the variational lower bound
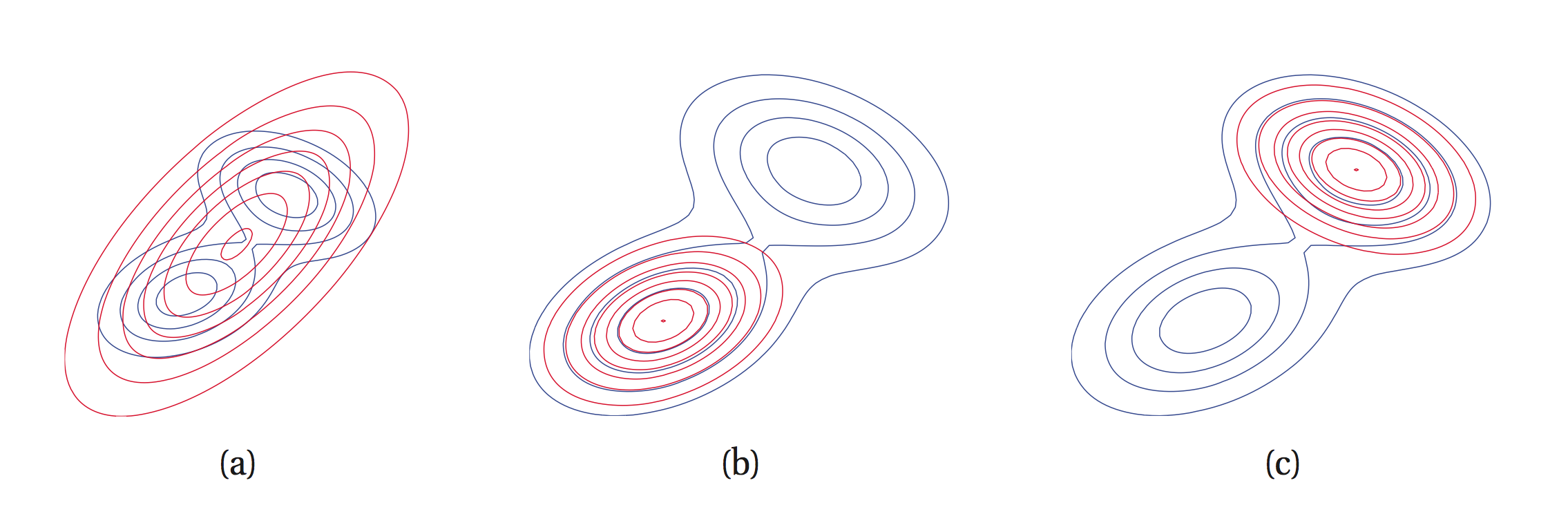
Now that we have our objective function, what distribution family is suitable to approximate a posterior density? (Or any density, for that matter?) Generally we want our distribution family to have nice properties, in that we want it to have closed-form (analytic) marginalisations, be smooth and continuous everywhere, infinitely differentiable, and computationally cheap to evaluate. For now let us assume that we use something like a normal distribution for our variational family. In principle now we have our objective function and our variational family, and there are many advantages (and some disadvantages) to using variational inference. Let's just summarise them so that you get a better picture of where we are going with this.
Advantages of variational inference
- Replace an expensive sampling operation with a cheap optimisation operation.
- Can approximate a posterior distribution very cheaply.
- Can interrogate (criticise) the model and update it with fast turnaround.
- Scales well to large data sets (that would be intractable).
- Enables things like Bayesian Neural Networks — with many layers — which gives our neural networks predictions with uncertainties, and uncertainty over our network architecture.
- Generalisable to many, many problems (any non-conjugate model).
Disadvantages of variational inference
- It's an approximate method: tends to under-estimate posterior variance and can introduce an optimisation bias.
- Non-convex optimisation procedure.
- Cannot capture correlations between model parameters.
- Posteriors are restricted in shape to that of the variational family.
For you, the biggest disadvantage to variational inference might be that you can't capture correlations between parameters in your posterior. Depending on your model parameterisation, that could be a big problem. There are many problems where we don't care about that though, including the entire field of mean field variational inference. Models like latent dirichlet allocation, or topic modelling, do not care about correlations between model parameters. We don't have time to talk about mean field variational inference here, so we are going to skip to doing automatic differentiation variational inference.
In automatic differentiation variational inference we are taking the variational family of latent variables and writing down the variational lower bound for the model, then using automatic differentiation during optimisation. If we can write down the variational lower bound then we can optimise the model and get closed form solutions for the approximate posterior of \(p\), in a fraction of the time it would take to sample. We still need to optimise this problem, and we can do this using stochastic optimisation (in TensorFlow, or any other automatic differentiation library), which allows us to scale our variational inference problem up to very large data sets.
The best variational inference package on the marketEdward, which was built on-top of TensorFlow. In Edward you only need to specify your model, and there are a number of examples already available. Disclaimer: Edward is to be integrated into TensorFlow, and the current Edward version is only compatible with an older version of TensorFlow (1.2), but you can see how this code ought to work. If you want to run your own example of variational inference, check out this notebook for PyMC3.
Let's start with a very simple example.
Here you can see that we have specified all of the network weights as TensorFlow variables. Then, instead of running any sampling, we use the ed.KLqp inference module which uses the K-L divergence as a metric for how close the posterior distribution and our approximate distribution need to be. That's it!
Here is an example for how we might construct a variational auto-encoder (using MNIST data) with Edward, allowing us to get approximate posteriors on the weights of the neural network! This example is derived from the Edward examples.
Summary
Machine learning — specifically neural networks — can be very useful for making predictions or by learning non-linear mappings from data. That's why they are universal interpolators. But frequently we also care about things that were not in the training set, and what predictions might arise for those kinds of objects. A standard neural network will always give us a prediction, even if the object was from outside the training set. And it can give us that prediction with a lot of confidence! Obviously this just tells ut that the notion of confidence in the output of a neural network is not akin to what we think when we think of probability. Instead it is a measure of relative belief that is conditioned upon the network, the training set, and a thousand decisions that you have made.
For all of these reasons we want our neural networks (in science) to have some degree of belief with what predictions they can make, and to recognise when there are inputs that are unusual from the training set. Bayesian Neural Networks are one way of doing this. Another, more flexible and powerful tool, is variational inference. Variational inference lets us turn a sampling problem (expensive) into an optimisation problem (cheap) by allowing for a family of distributions that can approximate the posterior. Variational inference can be used in lots of different fields of machine learning (see the Edward code example gallery) to make them efficient, or even tractable!
In our next (last) class, we will talk briefly about Recurrent Neural Networks and their applications within physics, before expanding on causality (or interpretability) in machine learning.